|
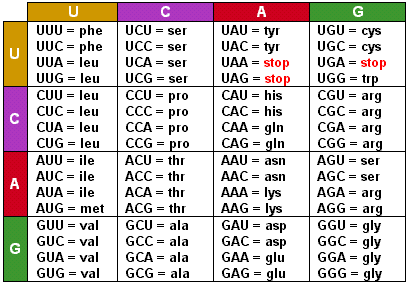
Answer the following questions about codons:ĪUG is unique because it serves as both the start codon, signaling the beginning of translation, and codes for the amino acid methionine. So, the amino acid sequence is: Met-Lys-Gly-His-ProĢ. Translate the mRNA sequence AUG AAA GGU CAC CCC into an amino acid sequence using the genetic code table. What would be the resulting amino acid sequence when your answer to 7a is translated?ġ. What would be the mRNA sequence when this mutated DNA is transcribed?ī. 5’ GGA GAT TAT 3’ (coding strand) 3’ CCT CTA ATA 5’ (template strand)Ī. Silent Mutations in DNA – Notice one nucleotide pair differs from the normal sequence given in question #6. What would be the normal amino acid sequence when your answer to 6a is translated? What would be the normal messenger RNA sequence when the template strand is transcribed?ī. Here is a normal (wildtype) DNA sequence: 5’ GGG GAT TAT 3’ (coding strand) 3’ CCG CTA ATA 5’ (template strand)Ī. The following problems examine some different types of DNA mutations and their effects. Mutations occur when there are changes in a DNA, RNA, or protein sequence. Many amino acids have more than one codon coding for them. Refer to the genetic code chart provided to answer the following questions. The genetic code consists of triplets of nucleotides called codons. See also the Structure Summary page (UniProt mapped Resources) for another way to access this information.Using the genetic code table provided, translate the following messenger RNA sequence into an amino acid sequence (protein). Clicking on that (see blue arrow) can open a page to display instances of polymer chains in the PDB that map to all or part of the UniProt sequence. However, in the row displaying the UniProt sequence, the UniProt ID for the sequence is included with a hyperlink. The Sequence Summary page is mostly used to integrate information about various aspects of the polymer being studied from various data resources. are located where the hydrophilic and hydrophobic regions are located where the mutations (if any) are present and much more.ĭepending on the polymer being displayed and the amount of information available about it, the sequence view pages can be very long and complex. Exploring this page can inform you about the structure and functions of the polymer - where the active site, binding site, etc. They help integrate information from a variety of resources and map them on the structure in an easily accessible format. The interactive sequence display provides a quick summary of the protein and nucleic acid polymers present in the structure. Are there parts of the structure that are poorly modeled or are missing from the coordinates?.Is the mutation at an important functional site (e.g., catalytic residue in an enzyme or a metal binding site)?.Does the polymer in the PDB have any mutations?.Is the polymer in the PDB entry the complete protein, just one or two domains, or merely a short peptide taken from the protein?.This display can be used to answer many questions, such as Viewing the sequence display can provide a quick way to assess how the polymer sequence of the PDB entry relates to the reference sequence (e.g., UniProt sequence). In addition, a variety of structural and functional annotations integrated from various sources (PDB, UniProt, and various bioinformatics resources) are also marked in this graphical display. A reference sequence from an external sequence database is displayed along with the polymer entity sequence where available. The protein and nucleic acid sequences of all entities are presented in the Sequence tab. In the PDB archive, an amino acid or nucleotide is usually represented by its one letter code using the FASTA format. The specific order of amino acids or DNA or RNA nucleotides in a polymer is referred to as its sequence. 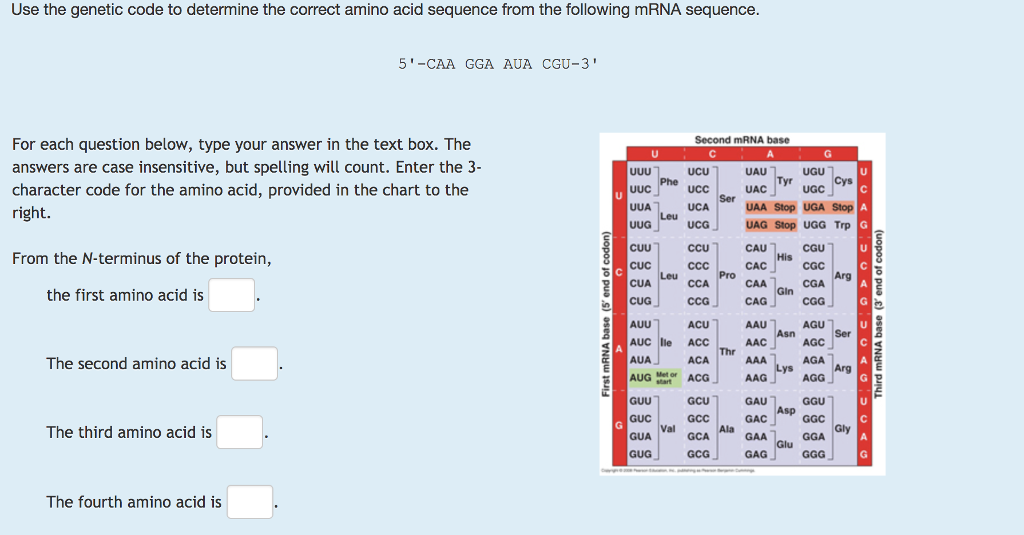
○ Exploring other structures Introduction What are sequences?
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
